The AGEM talent development grant 2020
Grant (€75.000) for exceptionally talented researchers who are in the first 5 years after obtaining a PhD-degree and want to develop their own research line. This grant has this year been awarded to:
Ruben Zapata-Perez
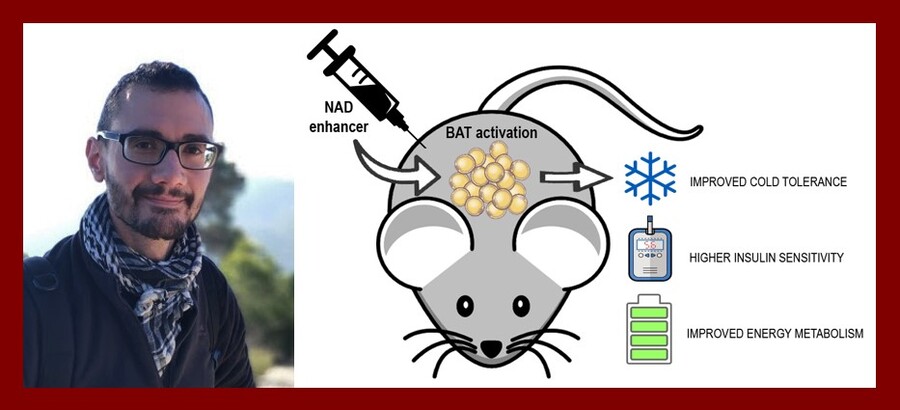
Enhancing brown adipose tissue metabolism through supplementation with novel NAD precursors
Brown adipose tissue (BAT) is a specific type of fat that helps maintaining body temperature by generating heat upon cold exposure. Due to its high metabolic activity, BAT plays an important role in the onset of cardiometabolic disease, a group of disorders comprising obesity, dyslipidemia and glucose intolerance. In fact, loss of BAT activity has been closely associated with high cardiovascular risk, while its activation is linked to the protection against atherosclerosis. Nicotinamide adenine dinucleotide (NAD+) can modulate BAT function through the activation of the NAD+-dependent deacetylases family of sirtuins, a family of proteins that tightly regulates cell energy metabolism. Activation of sirtuins by supplementation with NAD+ enhancers has proven to be an effective way to increase BAT activity in mice.
In the Laboratory Genetic Metabolic Diseases, we have identified two new compounds as highly active NAD+ enhancers. Following up on the promising results obtained in vitro, we now aim to validate their activity also in vivo. To do this, we will focus on the potential of these novel NAD+ enhancers to activate BAT function and improve the cardiometabolic disease outcome of high-fat-diet induced obese mice. This study will set the bases for the potential clinical application of these compounds.
The AGEM innovation grant 2020
Grant (€50.000) for innovative ideas beneficial to the AGEM research institute as a whole. This grant has this year been awarded to:
Chun-Xia Yi, Gertjan Kramer and Menno de Winther
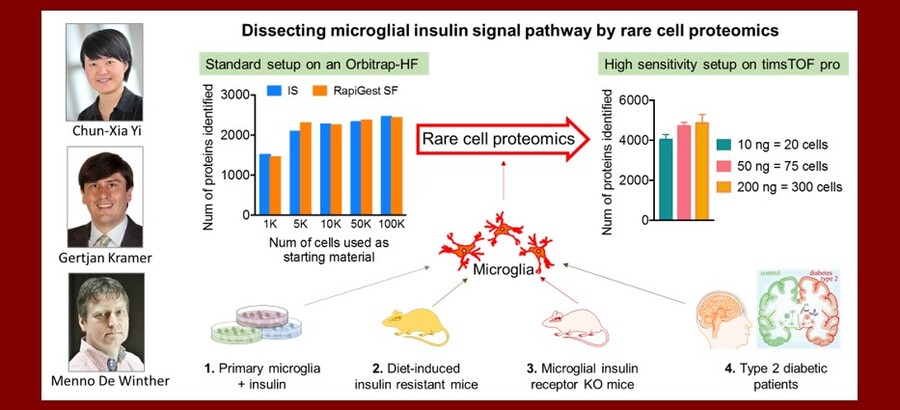
Dissecting microglial insulin signal pathway and insulin resistance by rare cell proteomics
Insulin resistance is a core feature of type 2 diabetes mellitus (T2DM). It takes place in both peripheral tissues and the brain. To understand how insulin signalling is impaired in the brain is pivotal for effectively treating T2DM and its associated brain diseases. So far, very little is known about insulin signalling and resistance in the brain innate immune microglia cells, that are responsible for maintaining a healthy microenvironment for neuronal survival and function. It is technically challenging to profile microglial protein expression, as the total number of microglia isolated from animals or human brain tissues is very limited. Therefore a state-of-the-art rare cell proteomics approach for deep proteome profiling is needed. This project will team up expertise from Dr. Chun-Xia Yi (Microglial pathophysiology in T2DM, Dept. Endocrinology and Metabolism, Amsterdam UMC), Dr. Getjan Kramer (Rare Cell Proteomics, Mass Spectrometry Core, SILS, UvA) and prof. Menno de Winther (immune cell metabolism, Dept. Medical Biochemistry, Amsterdam UMC), and to challenge the technical issue of dissecting insulin-signalling pathway and insulin resistance by rare cell proteomics in microglial cells.
Hilde Herrema and Peter Henneman
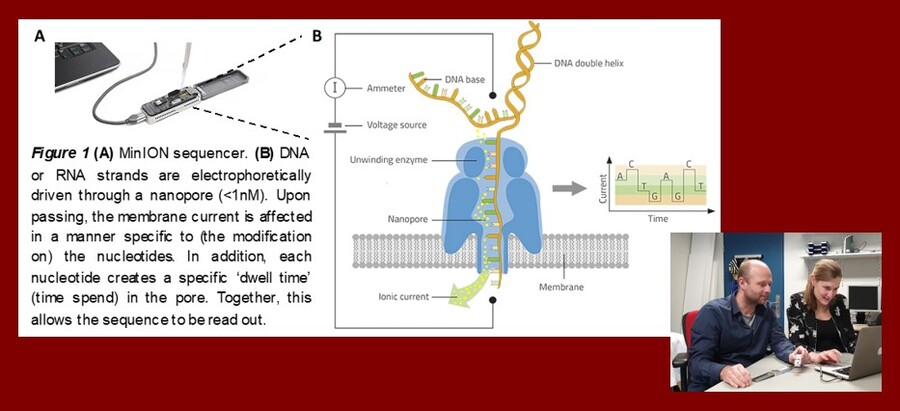
Nanopore sequencing; conductivity leads to connectivity
Life sciences have benefited tremendously from next generation sequencing technologies. Nevertheless, these technologies have proven to be suboptimal for particular applications. Third generation sequencers, such as the Oxford Nanopore long read sequencers, have the potential to significantly improve use of sequencing technologies for both research and diagnostic purposed. Nanopore allows direct sequencing of single DNA (or RNA) molecules and can create substantially longer reads (>100.000bp) than next generation sequencers (~300bp). Long read sequencing finds its application in de novo assembly of complex genomes (e.g. microbiome and phages) and can also be applied in the detection of large structural variation in the human genome (genome diagnostics). In addition, this technology is capable to detect and quantify modifications of DNA without a need for bias-inducing pretreatment (e.g., bisulfite conversion of DNA and PCR amplifications) of the sample. Using the AGEM Innovation Award, we aim to further expand our experience with long read sequencing. Moreover, we will make this knowledge and this exciting technology available for researchers within the AGEM and the Amsterdam UMC to further bridge initiatives at human and microbiota genomic translational research.
The AGEM matching grant 2020
Grant (€75.000 - €100.000) matched with an similar amount of money from (an)other source(s) for newly starting multidisciplinary research projects (PhD candidates or postdocs) that stimulate collaboration between (at least two) disciplines within the AGEM institute. This grant has this year been awarded to:
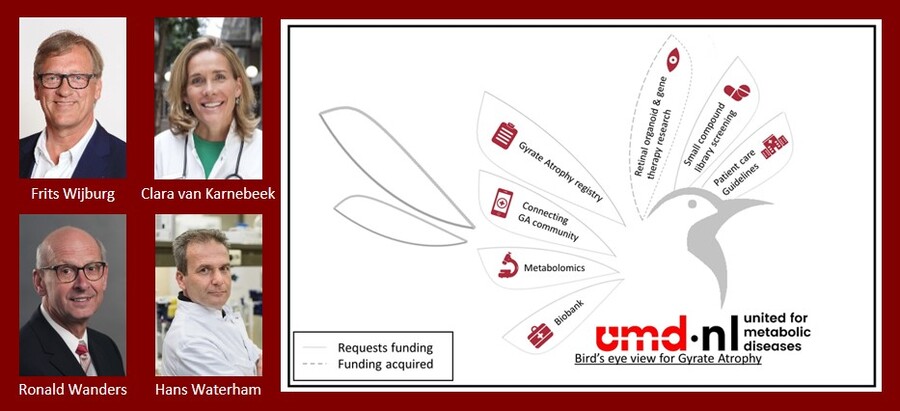
Hans Waterham, Ronald Wanders, Frits Wijburg and Clara van Karnebeek
Bird’s Eye View for Gyrate Atrophy
Gyrate Atrophy (GA) of the choroid and retina is an autosomal recessive metabolic disorder caused by a deficiency of ornithine δ-aminotransferase. Patients experience gradual reduction in peripheral vision during childhood followed by complete loss of vision around the 4th-5th decade. GA is rare and its true incidence is unknown and certainly underestimated. Scant literature is available, full clinical/biochemical phenotype and pathogenesis remain unclear, treatment options are limited and care often uncoordinated with poor outcomes. Our multi-disciplinary team’s mission is to prevent the debilitating blindness and other symptoms through this ‘Bird’s Eye View for Gyrate Atrophy’ project. With AGM funding we will establish a patient registry (clinical and biochemical phenotype, genotype, natural history, treatment outcomes), perform metabolomics analysis (pathophysiology, biomarkers), and high throughput compound screening (drugs increasing residual enzyme activity), increase awareness and patient-professional connections. This research will allow us to understand disease mechanisms, establish evidence-based recommendations for diagnosis and management, identify FDA approved compounds that increase residual OAT activity, and optimize trial-readiness also once gene therapy becomes available (via ongoing Amsterdam UMC organoid research). With these steps we aim to eventually diminish or even prevent the devastating blindness.
Jeroen den Dunnen, Riekelt Houtkooper, Marjolein van Egmond and Manon Wildenberg
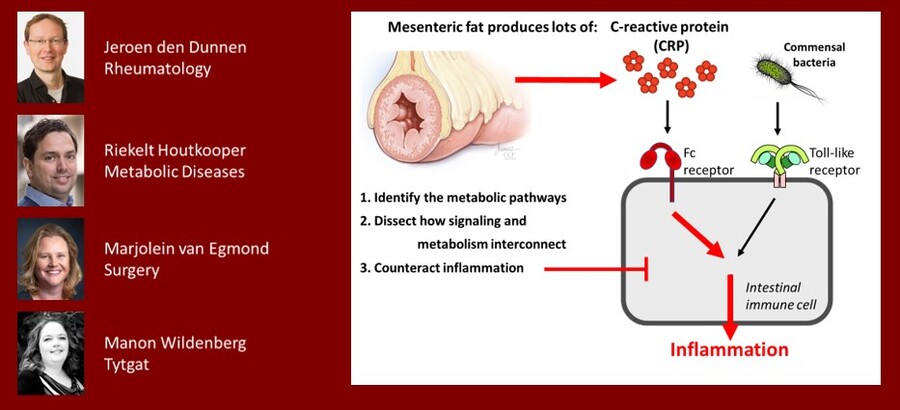
C-reactive protein: not only a marker, but also a cause of inflammation in Crohn’s Disease through metabolic reprogramming
One of the hallmarks of Crohn’s disease is the presence of mesenteric fat, which produces large amounts of C-reactive protein (CRP). Our preliminary data indicate that this CRP breaks the tolerance of intestinal immune cells for commensal bacteria, resulting in excessive inflammation. Interestingly, this inflammatory effect of CRP is fully dependent on metabolic changes inside the intestinal immune cells. In this study, four PI’s from different departments will team up to (1) identify this unique metabolic pathway, (2) unravel which signaling pathways activate the metabolic pathway, and (3) test therapeutic inhibitors to counteract the CRP-induced chronic inflammation.